There are a large number of abundant microbial resources in nature, but only 1% to 10% are currently cultivated and utilized by people, and a large number of microorganisms are not known and utilized by people. With the continuous development of genomics in the field of biotechnology, microbial genomics research has succeeded in the determination of the genome-wide sequence of Haemophilus influenzae by the genomic research (TIGR) using the shotgun method. In particular, by combining molecular biology research methods with microbiology research, people have made a series of important research results in the evolution, structure and genome of microorganisms, making it possible to utilize a large number of uncultured microorganisms. Among them, denaturing gradient gel electrophoresis (DGGE) technology is widely used in the study of microbial diversity. At present, DGGE has been applied to the analysis of bacteria, archaea, eukaryotes, micro eukaryotes and viral communities in the natural environment. Diversity. This technology provides superior species information in the community and analyzes multiple samples simultaneously, with repeatability and ease of operation. It is suitable for investigating the temporal and spatial changes of the population, and the community members can be identified by sequence analysis of the cut bands or hybridization analysis with specific probes. 3 DGGE application prospects Shine Skin Pumpkin Seeds,Shine Skin Pumpkin Seeds Does,Shine Skin Pumpkin Seeds Extract,Shine Skin Pumpkin Seeds Edible Wuyuan county dafeng oil food co.,ltd , https://www.dafengfood.com.cn
1 Basic principles of DGGE
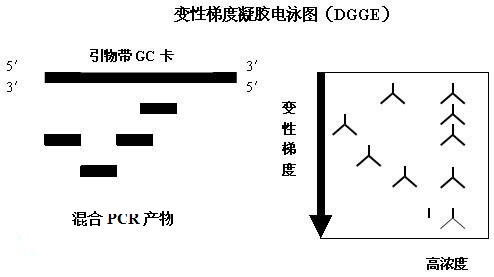
Figure 1 DGGE reaction principle In order to improve the detection sensitivity of DGGE, a GC-rich region (GC-Clamp, usually 30 to 50 nucleotides in length) is usually designed to be added to the 5' end of one primer. The complete melting of DNA increases the detection rate of different bases, so that sequences separated by one base can also be resolved.
2 Technical steps
The main operation process of this technology is as follows: (1) total DNA extraction of samples; (2) PCR amplification of 16SrDNA fragments of samples; (3) optimization and analysis of DGGE conditions; (4) subsequent processing such as tapping sequencing.
Since Handelsman et al. proposed metagenomics (Entagenomics) in 1998, the use of modern genomics techniques to study microbial communities in natural or artificial environments has become a hot trend. At present, metagenomics has been applied to the study of marine and soil microbial communities. Functional genes derived from uncultured microorganisms in the environment, and even genomics research, broaden the field of research on microbial communities. DGGE technology can monitor specific functional genes and their expression studies in complex environments, providing favorable information and genetic screening solutions for metagenomics research. In addition, the nucleic acid sequences obtained in DGGE analysis can be further applied to fluorescence in situ hybridization, DNA microarray and real-time PCR (PCR) to contribute to the rapid detection of specific populations.
DGGE is currently a widely accepted molecular biology tool for environmental microbiology complex and behavioral research in environmental microbiology. The technology is reliable, repeatable, fast and easy to operate. DGGE, which combines PCR amplification of marker genes or their transcripts (rRNA and mRNA), directly displays the dominant components of the microbial community. Because it can analyze multiple samples at the same time, it is very suitable for investigating the spatiotemporal changes of microbial communities, and it is possible to identify the members of the community by sequence analysis of the cut bands or by hybridization with unique probes. Coupled with the theoretical background of the role of this technology, such as the thermodynamics of the melting behavior of double-stranded DNA in solution. Use DGGE to understand the fate of a microbial community or some indicator microorganism after the environment is disturbed. Conventional use of functional genes as molecular markers can be used to identify closely related but ecologically distinct populations. End-labeled fluorescent PCR products and (intra-lane) standards can be used to detect rare population members and facilitate accurate comparisons between samples and samples. Double gradients (in combination with polyacrylamide gradients and denaturant gradients) can be obtained by DGGE analysis. Better resolution. In order to reduce the bias and limitations of various technologies, combined with PCR-DGGE and other molecular biology techniques and microbiological methods will more reveal the structure and function of microbial communities.
Although DGGE electrophoresis has many advantages in studying community dynamics and diversity, it does not provide information on metabolic activity, bacterial count, and gene expression levels. Therefore, it must be combined with other technologies to make up for the shortcomings. Both Biolog and enzymatic analysis can provide metabolic activity characteristics of the community. In addition, fluorescence in situ hybridization can provide relative information on bacteria, and in situ hybridization of stable isotope probes can provide metabolic information and obtain community structure and functional relationships. The 16S rDNA or gene library is also a method for analyzing the relative numbers of different populations. The number of bacteria and gene expression levels in the community can be quantified by real-time PCR amplification of 16S rRNA or functional genes of specific species. Therefore, combined with other molecular biology techniques, the efficacy of DGGE technology can be further enhanced to better serve microbial community structure and functional analysis. (Article Source: