Author: Nadia Tagnaouti1, Anitha Thomas1, Rebecca De Souza1, Ian Backstorm1, Andrew Brown1, Eric Ouellet1, Shyam Garg1, Grace Tharmarajah1, Keara Marshall1, Shannon Chang1, Timothy Leaver1, Andre Wild1, Oscar Seira2,3, Jie Liu2,3, Wolfram Tetzlaff2,3, Peter Deng4,5 , David J. Segal5 , Jan A. Nolta4 , Kyle D. Fink4 , R. James Taylor1 and Euan Ramsay1 An outer environment   1.NanoAssemblr TM Spark reactor 2. Pipette RNA and Neuro9 solution 3. Insert Spark, click "Start" 4. Aspirate and add to the cell 1. Prepare the nucleic acid solution to thaw the Neuro9 mixture into a separate syringe. Rat primary neurons were treated with Neuro9 GFP mRNA-LNP for 48 h in the presence of 5 μg / mL ApoE, and flow cytometry analysis of DIV 7 showed that > ​​95% of neurons absorbed LNP (calculated using nanoparticles) Fluorescent probes were administered and even at different therapeutic doses of GFP mRNA LNP (data not shown). Our results confirm that these lipid nanoparticles can be used as an efficient delivery system for CRISPR-Cas9 components to mediate genes. NanoAssemblr TM Spark Nanoparticle Synthesis System (Click here to open the product link) NanoAssemblr Benchtop Nanoparticle Synthesis System (Click here to open the product link) High Dispersion Homogenize Machine,Ultrasonic Homogenizer,Stomacher Blender,Handheld Ultrasonic Homogenizer Guangdong Widinlsa International Co.Ltd , https://www.gdwidinlsa.com
1 Precision NanoSystems Inc., Vancouver, BC, Canada, 2 International Collaboration on Repair Discoveries (ICORD), 3 Department of Zoology, 4 Stem Cell Program and Institute for Regenerative Cures, University of California Davis Health Systems, Sacramento, CA, USA, 5 Genome Center, MIND Institute, and Biochemistry and Molecular Medicine, University of California, Davis, CA, USA
In recent years, the demand for an efficient delivery tool capable of delivering payloads in vivo and in vitro to regulate gene expression has been steadily increasing. Lipid nanoparticles (LNPs) utilize a single-dose delivery pathway that binds apolipoprotein E (apoE) to mediate efficient delivery of encapsulated nucleic acids through the low density lipoprotein receptor (LDLR). However, their application from the experimental bench to the clinic has been considerably limited because of the challenges encountered in the manufacturing process. Here, we bridge this gap by describing the robust manufacturing and use of lipid nanoparticles. We use an optimized microfluidic platform to efficiently package and deliver nucleic acids (such as siRNA, mRNA, plasmid DNA) in a cell and animal model that is difficult to transfect in vitro at a manageable scale.
Here, we provide evidence for efficient cellular uptake of these LNPs in primary cortical rat neu- rons and their ability to deliver nucleic acid payloads that allow for down-regulation of targeted mRNA by siRNA-mediated degradation, expression of exogenous A sex mRNA sequence, as well as a gene that exogenously expresses a clone into a plasmid. These LNPs are capable of achieving high conversion efficiencies with no measurable associated toxicity. We also provided preliminary data detailing the efficiency of expressing genes in vivo using plasmid DNA-LNPs after striatum injection. In general, these studies were designed to efficiently combine small (siRNA) and large nucleic acids (mRNA, The strategy of delivering plasmid DNA to the original cells for treatment provides valuable insights. On this basis, we provide a suitable framework for efficient delivery of the RNA of the CRISPR-Cas9 system and expression of Cas9 by optimizing the LNP delivery method.
The delivery of genetic lipid nanoparticles (LNPs) via lipid nanoparticles is a platform that can be used to deliver nucleic acids to cells. LNPs mimic low-density lipoproteins (LDLs), which are taken up by endogenous pathways. LNPs are sensitive to pH and are designed to release their payload into the cytoplasm. 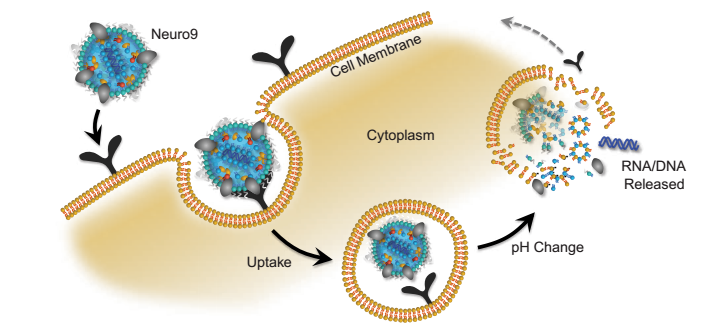
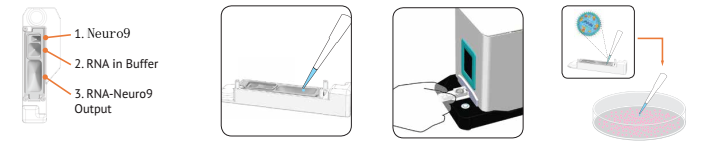
In vivo environment 
2. Place the syringe on the NanoAssemblr Benchtop and run the program for 3 minutes.
3. Centrifugal filtration or dialysis purification
4. Direct injection or systemic administration
Methods
1. Primary rat cortical neurons Cultured E18 rat cortical tissue was purchased from BrainBits, LLC. The cortex was removed from the transport medium and tryptic-stained trypsin-EDTA (ThermoFisher) with 0.25% tryptophan. The tissue was then washed with 10% FBS in DMEM (ThermoFisher) to inactivate trypsin-eda, followed by DMEM. Tissues were ground in NeuroCultTM Neuronal base Medium (StemCell) and NeuroCultTM, SM1 neuron complement (stem cells) and l-glutamine (stem cells) supplemented with 1-glutamate (Sigma) and then passed through 40 μm cells. A cell suspension filter is formed. The suspension was then counted and plated onto a PDL coated plate or glass at a density of 4.8 x 104 cells per square centimeter. Half of the medium was replaced with neuronal medium without L-glutamic acid every 3-4 days.
2. LNPs were treated with primary cortical neurons in rats 7 days later (DIV7) 5 μg / ml of ApoE was added at the indicated dose of LNP. Neurons were then incubated for 48 hours and harvested for endpoint assessment.
3. Flow cytometry After treatment and incubation with the indicated LNPs described above, neurons were washed with PBS and then separated from the culture dish by treatment with 0.25% trypsin-edta tryptophan. Inactivation of trypsin in PBS with 3% FBS, inactivating neurons to form a single cell suspension, then washing the neuronal pellets with PBS, resuspending in suspension with buffer (BD biosciences) and Apoptotic cells were stained with sodium iodide (BD biosceinces). Suspended stained neurons were then passed through a 35 μm cell strainer and then added using a BDCelesta flow cytometer.
4. Feasibility analysis According to the manufacturer's instructions, the standard protocol contained in the PrestoBlue® Cell Active Reagent (ThermoFisher) was calibrated to contain media tubes containing no neuronal media.
5. Immunocytochemistry In this evaluation, neurons were placed on a PDL-coated coverslip, then cultured, treated and incubated, washed with PBS, 4% PFA fixed neurons, with 0.1% trton - x Neurons were permeabilized and then blocked with NDS and then incubated with MAP2 antibody (Sigma M4403) for 1 hour. Neurons were then blocked in NGS and then incubated with GFP antibody (AbCam ab13970) overnight in NGS. A secondary antibody was then added and AlexaFluor-594 was used for MAP2 and alexafluor GFP-488. The coverslips were mounted on glass slides using an extended® diamond anti-fade patch (Thermofisher) and imaged on a confocal Microscope. It was confirmed by experiments that the selective binding of the first and second antibodies has no main control (data not shown)
6. RNA extraction and RT-qPCR
After processing and incubating the neurons, pre-form RNA extraction (hermoFishe) using the PureLink® RNA Mini Kit RNA Extraction Kit according to the manufacturer's instructions. RNA concentration determination was performed using NanoDrop (ThermoFisher) and then the same amount of RNA was used for cDNA conversion using the SuperScript IV VILO Master Mix (ThermoFisher) instructions. The target mRNA and ActB obtained from IDT were subjected to RT-qPCR using an TaqMan primer and probe, iTaq Universal Probes Supermix master mix (BioRad), and a thermal cycle (BioRad) was run on BioRad CFX96, three replicates per PCR. The expression value of the target mRNA was normalized to the ActB value using the ∆∆Ct method.
Transmission of mRNA in rat primary neurons 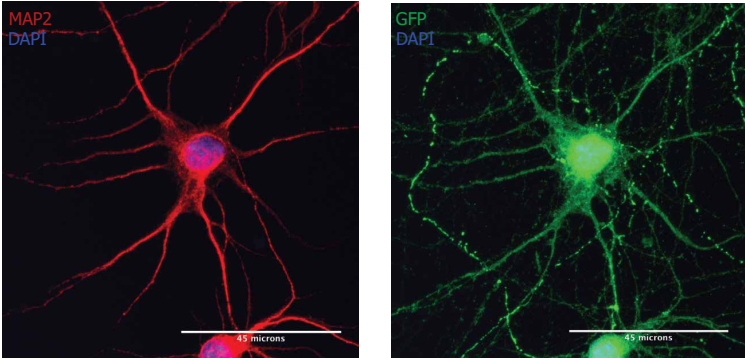
Note: MAP2-positive neurons (red) were treated with Neuro9 GFP mRNA-LNP 5 μg/ml ApoE to express GFP (green). Cellular anti-map2 primary antibody/alexa fluor-594 conjugated secondary antibody and anti-GFP primary antibody/alexa Fluor-488 coupled secondary antibody staining. Nuclear staining with DAPI (blue) 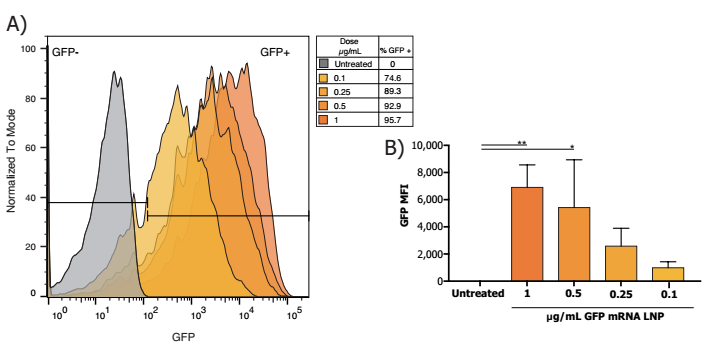
Delivery of plasmids in rat primary neurons 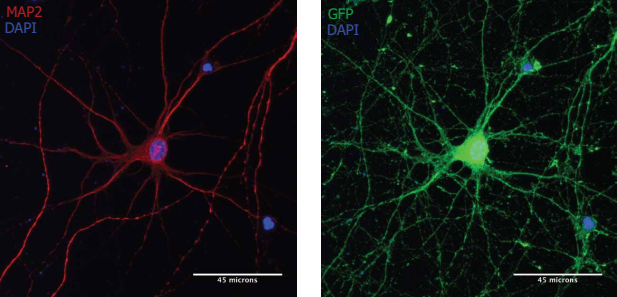
MAP2-positive neurons (red) were treated with Neuro9 GFP to express GFP (green) plasmid-LNP 5 μg / ml ApoE, cells were conjugated with anti-map2 primary antibody/alexa fluor-594 and anti-gfp primary antibody/alexa fluor -488 coupled secondary antibody staining, nuclear staining with DAPI (blue) 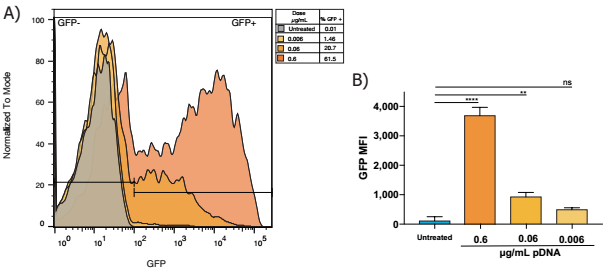
Flow cytometry analysis in the presence of Neuro9 GFP plasmid-LNP for 48 h after treatment 5 μg / ml ApoE in murine primary neurons, DIV 7, showed > 95% of neurons showed lnp absorption (calculated with fluorescent probes) , there are indeed packages in the nanoparticles) treated with different doses of GFP plasmid (data not shown) 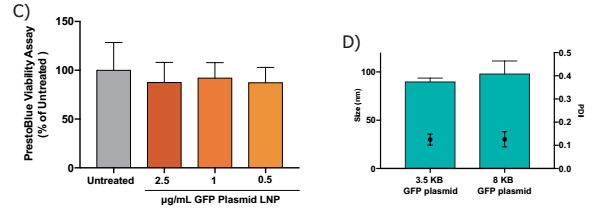
Delivery of plasmids in vivo 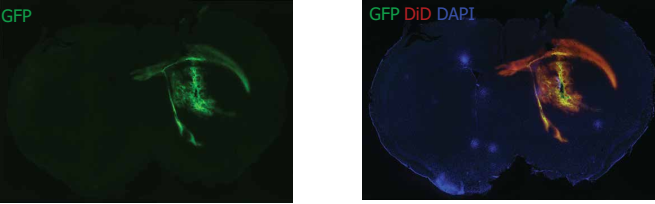
FVB mice at 10 months of age were injected subcutaneously with GFP plasmid-LNP and after GFP expression at 48H (green). 5 μl of 3 mg/ml GFP plasmid-LNP was injected. The nuclei were stained (blue) with DAPI, LNP stained with red (red), and GFP expressed (green) on the same region. Both LNP uptake and GFP expression localize the striatum within the injection area and have spread to peripheral proliferative regional systems and corpus callos including the ventricular portion.
Small RNA transfer in rat primary neurons 
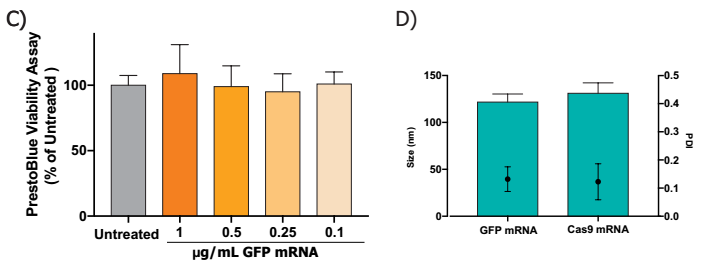
B) Cell viability was measured using a PrestoBlue® Cell Viability Reagent. At an effective dose of 5 fold, no toxicity was observed, using one-way ANOVA and Dunnett's multiple comparison test. 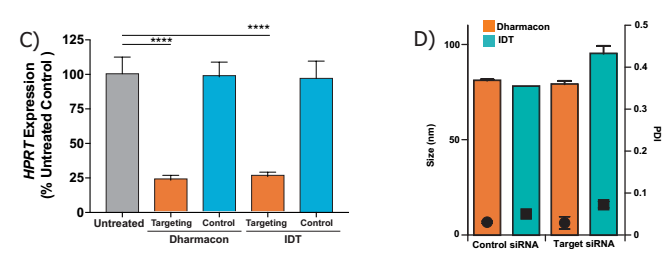
C) siRNA LNP dose at 1 μg / ml of product, using both IDT and Dharmacon siRNAs, targeted mRNA 75% knockdown, n=3 per treatment, and Dunnett's one-way ANOVA multiple comparison test, **** p <0.0005.
D) C) Comparison of the size of the nanoparticles encapsulating different siRNAs with PDI, although using different sizes of siRNAs, there was no significant difference in particle size and PDI.
in conclusion
LNP-mediated CRISPR-Cas9 Delivery
LNPs have multiple functions to deliver Cas via mRNA, pDNA or pre-formed ribosomal protein complexes, and one or more targeting strands are also well delivered. 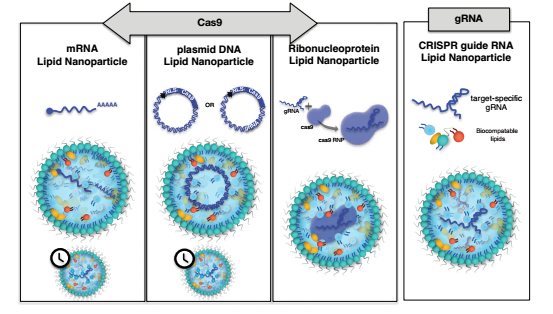
laboratory apparatus: 
