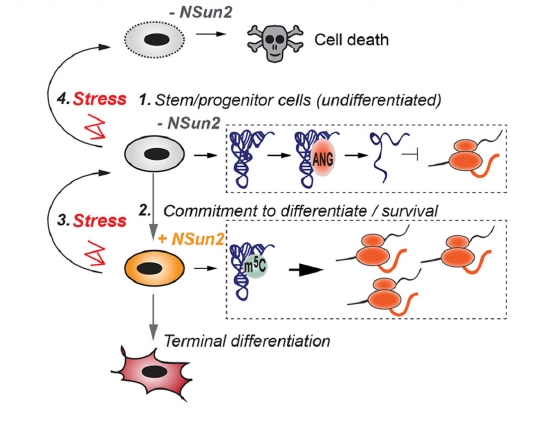
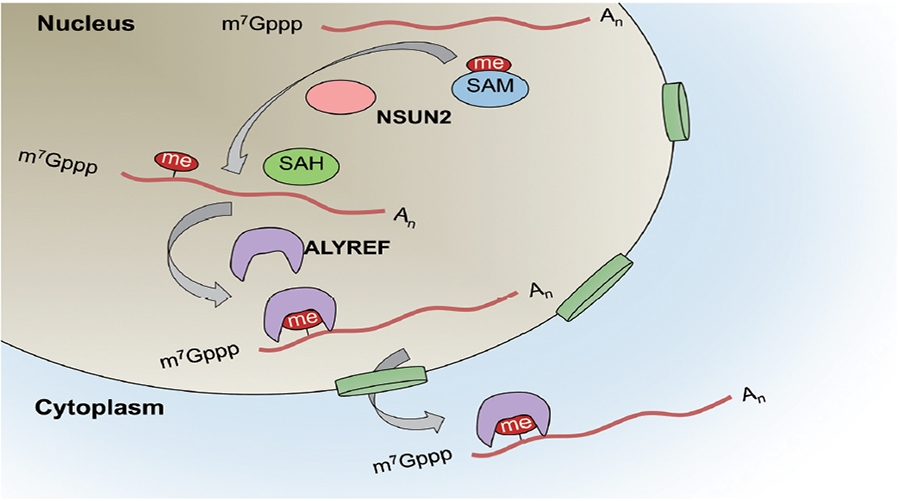
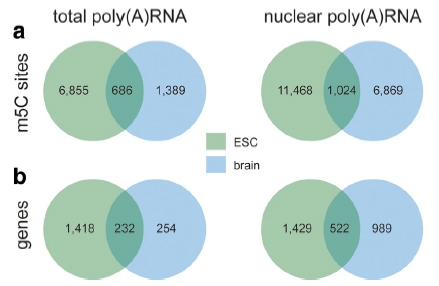
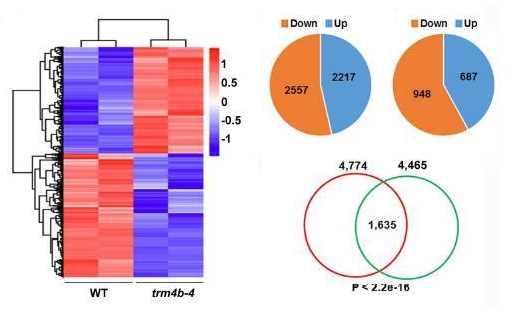
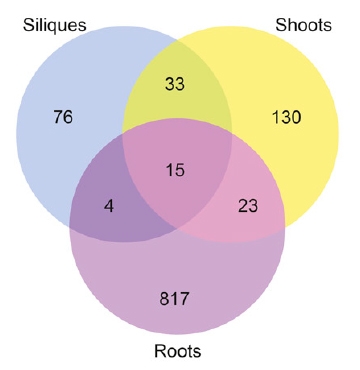
RNA methylation is a research hotspot in the field of epigenetics and the darling of Nature, Science-level articles. With the deepening of RNA methylation research, a new class of RNA methylation modification has once again set off a hot topic in the scientific community. It is a 5-methylcytosine (m5C) RNA methylation modification in tRNA and rRNA. Medium and high abundance is stable. Its function has been shown to regulate stem cell stress, cytotoxic stress, mRNA nucleation and plant cell development and gene expression. As a pioneer in RNA methylation research, Yunzoo is the only company on the market that provides m5C RNA methylation sequencing services. Today we undertake the last m6A methylation and continue to bring you the latest research summary of m5C methylation. 1. Nature: m5C regulates stem cell function and stress stress Impact factor: 40.14 Protein synthesis is a fundamental process in all cells, but its precise regulation in development, stem cells and cancer is unclear. Using m5C RNA methylation sequencing technology, the team found that post-transcriptional methylation of NSun2 on cytosine-5 (m5C) is a novel mechanism for inhibition of total protein synthesis, thereby confirming m5C RNA methylation as a An important pathway to regulate total protein synthesis and cell fate. NSun2-mediated methylation protects tRNA from being cleaved into non-coding 5' tRNA fragments, thereby facilitating protein translation and differentiation. External stress stimulation inhibits the activity of NSun2, causing it to cleave into 5' tRNA fragments and then reduce protein synthesis in human cells. Inhibition of post-transcriptional methylation in squamous tumors promotes stem cell function and tumorigenesis. However, cytotoxic stress requires reactivation of the m5C RNA methylation pathway to exit a specific translational repression program. Therefore, activation of RNA methylation or inhibition of tRNA cleavage is critical for tumor-initiating cells to respond to the survival of cytotoxic stress. Figure 1. m5C RNA methylation regulates stem cell function 2.Cell Research: m5C regulates mRNA export Impact factor: 14.81 The research team first used m5C RNA methylation sequencing and bioinformatics analysis technology to reveal the distribution of mRNA m5C, and mapped the m5C modification map. It was found that m5C was mainly distributed in CG-rich regions. By analyzing and comparing different tissues of human and mouse, it was found that the distribution characteristics of m5C on mRNA were very conserved in mammals, while the genes modified in different tissues were specific. The study found that NSUN2 (Writer) protein is the main mRNA m5C methyltransferase, its activity depends on C271 and C321 sites, and the loss of NSUN2 function leads to inhibition of mRNA nucleation. Further studies indicated that the nuclear regulatory protein ALYREF (Reader) specifically binds to the m5C modification site by the 171th lysine, thereby promoting mRNA nucleation. Figure 2. m5C regulates mRNA nucleation 3. Genome Biology: M5C modification is widely present in mouse embryonic stem cells and brain tissue Impact factor: 11.91 The research team at Innsbruck Medical University used m5C RNA methylation sequencing and fluorescence quantification to find that there are a large number of m5C methylation sites in mRNA and non-coding RNA, and there are large differences in the number and distribution of different samples. . Compared with brain tissue, the diversity of methylated mRNA in ESCs (embryonic stem cells) is higher. GO analysis indicated that methylation is enriched in highly proliferating ESCs and is predicted to be associated with cell cycle, RNA and chromatin modifications; whereas in brain tissue, methylated transcripts are involved in ion transport and synaptic function Enrichment in categories. Particularly in ESCs, most sites that are methylated in ESCs are not methylated in brain tissue samples. Therefore, the researchers believe that differential methylation of RNA, which may be different cell types, is involved in regulating the transcription and translational properties of specific RNAs. Figure 3. Comparison of mouse methylation of mouse ESCs and brain tissue m5C 4. Molecular Plant: Arabidopsis m5C RNA methylation profile Impact factor: 9.33 The research team of the Chinese Academy of Agricultural Sciences, Gu Xiaofeng, explored the new mechanism of plant RNA methylation regulation by exploring the methylation of m5C RNA in Arabidopsis thaliana. The researchers used RIP sequencing to reveal the distribution of 6045 m5C methylation enrichment regions on Arabidopsis mRNA. At the same time, m5C methyltransferase was discovered and a new mechanism for regulating plant development and gene expression was described. This study reveals that m5C is a new apparent regulatory marker in plant genes and is involved in the regulation of plant growth and development. The research results provide a basis for further exploration of RNA methylation regulation mechanisms and applications. Figure 4. Up-regulation and down-regulation of RNA-seq validation genes 5.The Plant Cell: Arabidopsis mRNA and non-coding RNA m5C methylation profile Impact factor: 8.23 The team used m5C RNA methylation sequencing to quantify the transcriptome of m5C methylation in Arabidopsis, resulting in mRNA, lncRNA and other ncRNAs in Arabidopsis three tissue types (fruit, shoot and root) More than 1000 m5C modification sites were found. Quantitative differences in methylation sites between the three tissues suggest tissue specificity for m5C modification. Interfering RNA m5C methyltransferase TRM4B results in loss of m5C sites on mRNA and ncRNA and decreases the stability of tRNAAsp (GTC). The researchers also demonstrated the importance of m5C in plant development because the trm4b mutant has a shorter primary root than the wild type due to reduced cell division in the apical meristem. In addition, the trm4b mutant showed increased sensitivity to oxidative stress. Figure 5. Detection of m5C modification levels in Arabidopsis using bsRNA-Seq Cloud order related product recommendation: m6ARNA methylation sequencing m5CRNA methylation sequencing m1ARNA methylation sequencing RIP sequencing RNA Pull Down Past review The latest cloud sequence biology m6A "RNA methylation" research summary - cancer articles The first domestic 10 points or more m6A RNA methylation sequencing article - cloud sequence bio-help! Cloud order customer 12 points top article, teach you how to use RIP sequencing to play the molecular mechanism! Yunxu Bio exclusive M5C RNA methylation sequencing 2018 National Nature Research Hotspot 2: In-depth analysis of RNA methylation research RNA methylation in 20 points, coexistence of heat and strength Genome research: There are huge differences in methylation modification of different RNA m5C Plant Cell: Arabidopsis found a new m6A RNA demethylation modifying enzyme Molecular cell: a new method for m1A RNA methylation sequencing--m1A-MAP Hepatology: m6A RNA methylase METTL3 promotes the development of liver cancer Nature Heavy: m6A RNA methylation is involved in the key aspects of hematopoietic stem cell development Cancer cell: Professor He Chuan discovered the important function of m6A RNA methylation Genome Biology: Using RNA sequencing to publish RNA methylation 10 points RNA methylation book (2) m6A methylation and MeRIP sequencing m5C RNA methylation research method Shanghai Yunxu Biological Technology Co., Ltd. Shanghai Cloud-seq Biotech Co.,Ltd Address: Lane 1066, Qinzhou North Road, Caohejing High-tech Development Zone, Shanghai phone Website: mailbox: Sports Towel,High Quality Towel Robe,Quick Dry Sport Towel Robe,Sport Towel Supplier Suzhou Golden Gamrnet MFG Co.,Ltd , https://www.suzhoumfg.com